Distinct differences in the tumour-resident microbiome between early- and late-onset colorectal cancer
Abstract
Background: The incidence of early-onset colorectal cancer (EOCRC; under 50 years of age) is rising globally. Changing environmental exposures and their influence on the colonic microbiome is thought to be a key factor in this increasing incidence. While differences in the gut microbiome have been reported between EOCRC and late-onset CRC (LOCRC) patients, little is known about differences in the tumour-resident microbiome. This study aims to characterise the differences in the tumour microbiomes between EOCRC patients and older CRC patients.
Methods: RNA sequencing was carried out on pre-treatment tumour samples from a cohort of EOCRC patients. Meta-transcriptomic data was compared to samples from a control group of LOCRC patients aged over 65 years. We compared the microbial alpha and beta diversity as well as differential abundance of microbial taxa between EOCRC patients and the older controls.
Results: There were 19 samples in the EOCRC group, compared to 196 LOCRC samples. Tumours from EOCRC patients had decreased observed species richness (p=0.009), but no significant difference seen in Shannon alpha diversity (evenness and richness) (p=0.16) or beta diversity, compared to LOCRC tumours. We identified two microbial phyla, 14 genera, and 18 species that were significantly enriched in EOCRC tumours compared to LOCRC tumours (p=<0.05).
Conclusion: The tumour-resident microbiome of EOCRC is distinct from that seen in older patients. Changing environmental exposures and the impact that these have on patients’ microbiomes may play a role in driving the increased incidence of EOCRC.
Introduction
The incidence of early-onset colorectal cancer (EOCRC) has been rising in many countries, globally, including Germany, the United Kingdom, Denmark and Aotearoa New Zealand.1,2 The greatest increases in incidence have been seen in distal colonic and rectal cancers.3 Some studies have estimated that if current trends continue, nearly one in four rectal cancers will be diagnosed under the age of 50 years, by 2030.4
There are distinct clinical characteristics between EOCRC and late-onset colorectal cancer (LOCRC). EOCRC patients are more likely to present with later-stage disease (stage 3 or 4), 5 and have higher rates of mucinous or signet ring histology and poorly differentiated cancers.6 Research has also shown that EOCRC responds differently to treatment compared with LOCRC patients, and therefore may have a distinct underlying biological behaviour.7
The recent epidemiological trend of increasing EOCRC incidence is occurring independently of trends in older adults; in many countries, the increasing EOCRC incidence is occurring in a background of declining overall CRC incidence, implying that changing environmental exposures, unique to younger generations, may be causative.3,8 The changing environmental ‘exposome’9 and early-life exposures can impact the colonic microbiome leading to microbial dysbiosis and a carcinogenic environment.10 An example of this is how an increasingly ‘Western’ diet, including more fatty, meaty, processed and sugary food, impacts the gut microbiome by causing a shift in dominant microbiota and their metabolic activity, resulting in increased sulphur metabolism, which is associated with the development of EOCRC.11 Likewise, other factors, including increasingly sedentary lifestyles, alcohol intake, obesity, caesarean section rates and increased antibiotic usage, can adversely impact the intestinal microbiome, which acts as the interface between environmental stressors and human homeostasis; thus, shifts in the gut microbiome may drive increasing incidence of EOCRC and may also influence the tumour-resident microbiome in CRC and activation of associated molecular pathways in the tumour microenvironment.10
Several bacterial genera are associated with the development of CRC, including Bacteroides, Peptostreptococcus, Streptococcus, Prevotella, and Escherichia.12, 13, 14 Conversely, other taxa, such as Lactobacillus15 and Bifidobacterium16 track with reduced CRC risk. Previous studies have demonstrated differences in both the faecal17 and tumour18 microbiomes between EOCRC and older CRC patients. However, the mechanisms by which the microbiome and how specific bacteria may play a part in the rising incidence of EOCRC is poorly understood. Thus, there remain significant gaps in our knowledge about their role in driving EOCRC.
This study aims to describe differences in the microbial transcriptome between tumour samples from a cohort of EOCRC and LOCRC. We will also discuss our current understanding of how the microbiome may contribute to colorectal carcinogenesis, and how this may be relevant to the development of EOCRC.
Methods
Patients and Samples
Two cohorts were used in this study. The first cohort19 consisted of 260 surgical resections of colorectal tumours from patients at Christchurch Hospital, Aotearoa New Zealand. Samples were taken during surgery, frozen in liquid nitrogen, and stored at -80°C prior to RNA extraction. Exclusion criteria for this cohort included chemotherapy/radiotherapy prior to the study, diagnosis of Hereditary Non-Polyposis Colorectal Cancer (HNPCC) or Familial Adenomatous Polyposis (FAP). All participants in this cohort provided written, and informed consent. The study was approved by the University of Otago Human Ethics Committee (approval number: H16/037).
The second cohort20 consisted of tumour biopsies from 40 rectal cancer patients treated at Christchurch Hospital, Aotearoa New Zealand and the Peter MacCallum Cancer Centre, Melbourne, Australia. Tumour biopsies were taken at colonoscopy and immediately frozen in liquid nitrogen, and stored at -80°C prior to RNA extraction. Exclusion criteria included previous chemotherapy or radiation therapy. All participants provided written, informed consent. Ethical approval for this cohort was given by the Health and Disability Ethics Committee, New Zealand (approval number: 18/STH/40/AM01), and the Human Research Ethics Committees of Australia (approval number: HREC 14/85).
We performed analyses comparing the differences between two different age groups. The first group consisting of all those aged 50 years or younger, termed early onset (EO), with a control group consisted of all those aged over 65 years, termed late onset (LO). We excluded those patients aged between 50-65 years in order to better delineate differences between early- and late-onset CRC cohorts.
RNA extraction, Sequencing and Processing
We extracted RNA as detailed previously.21 Briefly, we used the RNEasy Plus Mini Kit (Qiagen) to extract RNA from 15-20 mg of sample. The tissues were first disrupted using a Retsch Mixer Mill, and a DNAse treatment step was included in the procedure. Purified RNA was quantified using a NanoDrop 2000c spectrophotometer (Thermo Scientific, Asheville, NC, USA) and subsequently stored in −80 °C.
RNA Sequencing, library preparation, and host gene and microbiome taxa profiling were performed as detailed in previous studies.22,23 Briefly, we used MetaFunc24 for read processing; we mapped reads that have undergone quality control to the human genome, and unmapped reads were run through Kaiju25 to identify microbial taxonomies.
Microbial Analyses
Diversity
We rarefied the sample set to 90% of the smallest sample size in the dataset, and obtained alpha diversity measures per sample using Phyloseq’s26 estimate_richness function for Observed (richness) and Shannon (richness and evenness) measures, and compared differences in the alpha diversity measures between EOCRC and LOCRC groups using the Wilcoxon test built into ggplot2’s built-in stat_compare_means() function.
We visualized microbiome community differences between groups using Phyloseq’s ordinate function on a center-log-ratio-transformed dataset, using RDA (redundancy analysis) as method and euclidean distance.
Microbial Taxa and Differential Abundance Analyses
We compared phyla, genus, and species levels of microbial taxonomies between our two groups. For phylum level analysis, agglomerated identification to the phylum level, obtained percent abundance for each phyla, and compared relevant taxonomies using Wilcoxon tests. We used DESeq227 with zero-inflation modelling using zinb-wave28 to compare microbial abundances at the genus and species levels between EOCRC and LOCRC groups, accounting for sequencing batch, sex, and location covariates. We considered a microbe differentially abundant if the adjusted p-value is <0.05.
Results
The meta-transcriptomes of CRC tumour tissue from a total of 300 patients were analysed. The cohort comprised of 19 patients aged 50 years or younger (which was termed early-onset), and 196 aged over 65 years (termed late-onset), which was used as a control group. Eighty-five patients between the ages of 50 and 65 years were excluded from our analyses (Table 1)
The proportion of sexes between the two groups was comparable, with the early-onset group being 53% female, vs 55% in the late-onset group. There was a higher proportion of rectal cancers in the early-onset group (47%) compared to the late-onset group (26%), however due to our small sample size this result did not reach significance (p=0.06) (Table 1).
Table 1: Cohort demographics
| Early onset | Late onset | p-value | |
|---|---|---|---|
| n (%) | n (%) | ||
| Total | 19 | 196 | |
| Sex | |||
| Female | 10 (53%) | 108 (55%) | |
| Male | 9 (47%) | 88 (45%) | 1.00 |
| Location of Tumor | |||
| Colon | 10 (53%) | 145 (74%) | |
| Rectum | 9 (47%) | 51 (26%) | 0.06 |
Microbial Diversity
Alpha diversity refers to community diversity in a sample, defined by how many different taxonomies are in a sample and how different the abundances of each taxonomy are between conditions. In this study we measure both observed alpha diversity (species richness) and Shannon alpha diversity (species richness and eveness). There was a significant difference in observed alpha diversity seen, with decreased observed species richness in EOCRC compared to LOCRC (p=0.0087) (Figure 1A). However, there was no significant difference seen in Shannon alpha diversity (evenness and richness) (p=0.16) (Figure 1B). Beta diversity describes how different microbial communities are between two groups. We did not observe any significant separation between the two groups (Figure 1C).
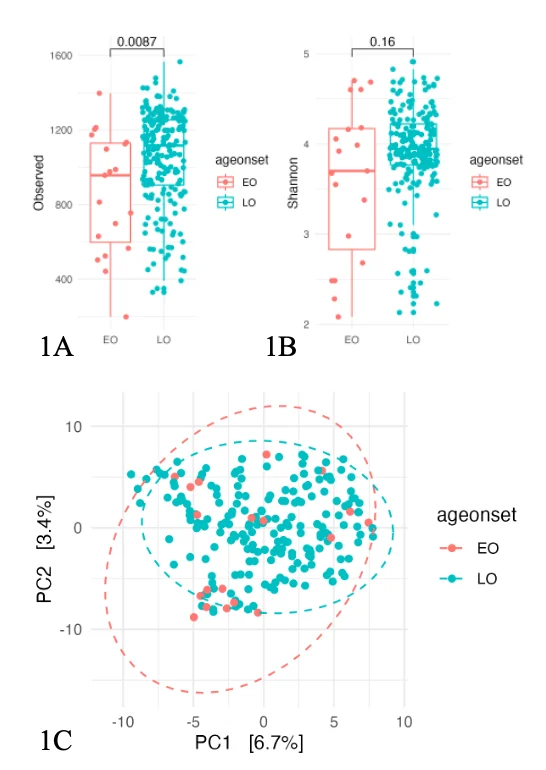
Figure 1. Diversity analysis between early-onset and late-onset cohorts. 1A shows Observed alpha diversity (p=0.0087), 1B shows Shannon alpha diversity (p=0.16) and 1C depicts beta diversity. EO, early-onset; LO, late-onset; P-value <0.05 = significant.
Phylum-level analysis
We then compared the abundance of the top phyla between the two cohorts. Any phyla with a median relative abundance <1% was excluded. Of the top five phyla, there were two that were significantly different between the two groups, with significantly lower abundance of both Chlamydiae (p-value = 0.018) and Actinobacteria (p-value = 0.036) in the early-onset cohort. There was no significant difference in abundance of Proteobacteria, Bacteriodetes or Firmicutes (p-value > 0.05) (Figure 2).
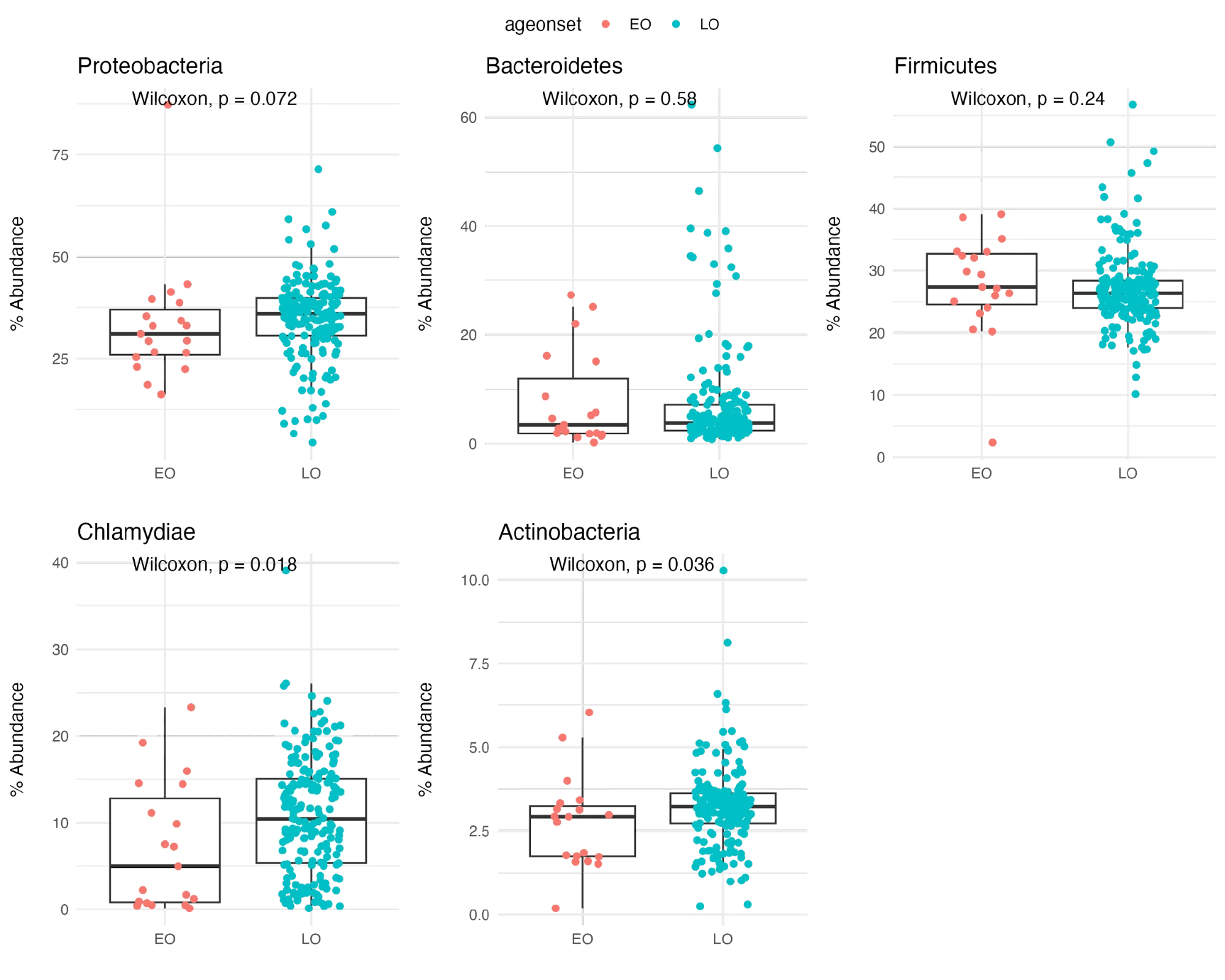
Figure 2. Differences in abundance of top phyla between early-onset and late-onset cohorts. EO, early-onset; LO, late-onset; P-value <0.05 = significant.
The Firmicutes/Bacteroidetes ratio, a commonly used measure of adverse microbiome changes, was measured in the two groups and were found to be comparable, with the ratio being 10.04 in the early-onset and 8.06 in the late-onset group (p-value = 0.52).
Microbial Genus-level Analysis
Fourteen microbial genera were found to be significantly more abundant in the EOCRC samples, including Anabaena and Schaalia. Four genera were more abundant in the LOCRC cohort (Figure 3).
We also looked specifially at the abundance of selected bacterial genera previously associated with CRC, including Fusobacterium, Bacteroides, Porphyromonas, Prevotella, Peptostreptococcus and Parvimonas, and found no significant difference in abundance in these between the two patient cohorts.
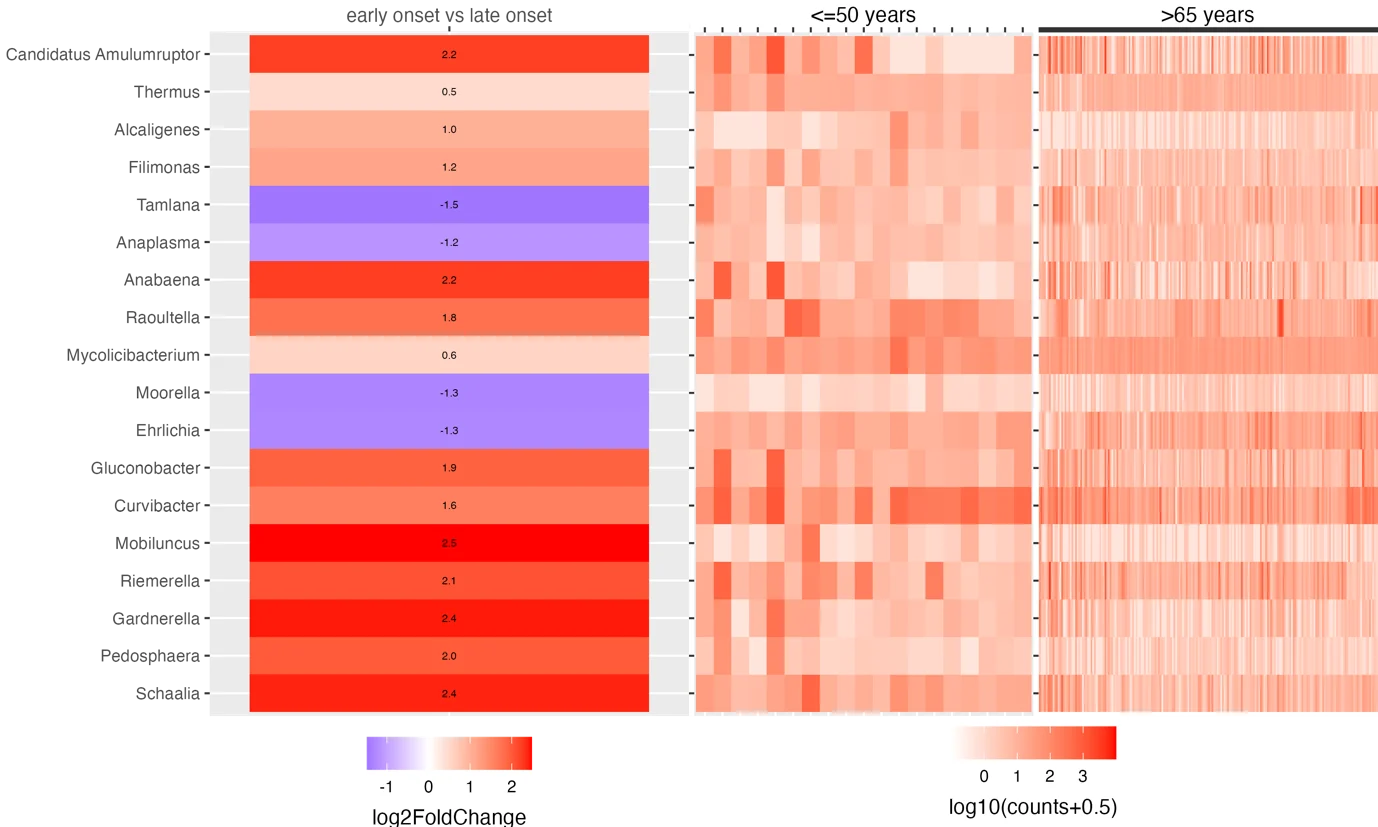
Figure 4. Microbial genera found to be differentially enriched in early-onset and late-onset tumours. Heat map showing the microbial genera that were differentially enriched between early-onset and late-onset (p.adjust <0.05, and log2FoldChange >0 for early-onset, and <0 for late-onset).
Microbial Species
Eighteen bacterial species were found to be significantly enriched in tumours from the early-onset cohort, compared to late-onset patients. These included two species of Bacteroides and one species of Klebsiella. Two bacterial species, Campylobacter ureolyticus and Helicobacter canadensis were enriched in tumours from the late-onset group, compared to early-onset patients (Figure 4).
We further anaysed differences in abundance of several microbial species, also known to be associated with the developement of CRC, including Fusobacterium nucleatum, Bacteroides fragilis, Bacteroides thetaiotaomicron, Porphyromonas assacharolytica, Parvimonas micra, and Prevotella intermedia between the two cohorts and found no statistically significant difference between the two groups in the abundance of these species.
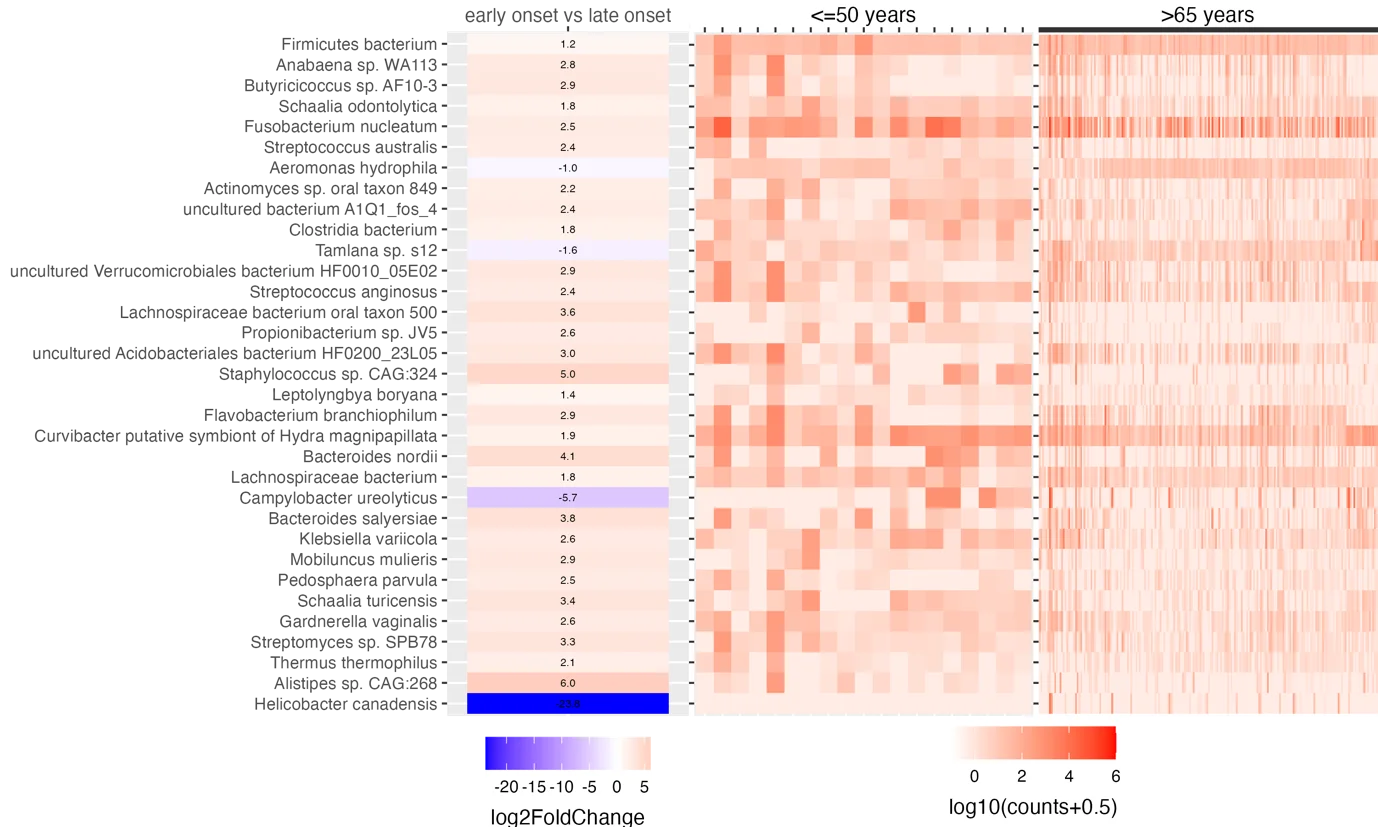
Figure 5. Microbial species found to be enriched in the early- and late-onset tumours. Heat map showing the microbial species that were differentially enriched between early-onset and late-onset (p.adjust <0.05, and log2FoldChange >0 for early-onset, and <0 for late-onset).
Discussion
In this study, we have investigated the differences in tumour microbial diversity, and abundance of bacterial phyla, genera and species between early-onset and late-onset colorectal tumours. While differences in the microbiome have been implicated in the development of CRC in general for some years, there is little information available regarding specific patterns in the tumour microbiome that may be driving the development of early-onset colorectal cancer, and how differences in the tumour microbiome between the two groups could help to explain the differences in clinical and molecular characteristics between the two groups. Our results suggest that patients with EOCRC have unique tumour-resident microbiomes compared to patients with LOCRC, with altered diversity and different patterns of enriched microbial phyla, genera, and species.
There is evidence that changes in the microbial environment may contribute to the different molecular and clinical characteristics of EOCRC, compared to late-onset disease. Research by Flynn et al,29 found that the microbiome of the mucosa and the lumen of the healthy colon is different in the proximal colon compared to the distal colon and rectum, suggesting a potential link between these differences in the gut microbiome and the patterns of tumour location seen. Likewise, specific tumour-resident bacterial profiles have been linked to different CRC consensus molecular subtypes (CMS) of CRC, for example, enrichment of Bacteroidetes has been associated with CMS 1 tumours, while enrichment of Prevotella has been associated with CMS 2 subtypes.21 Analyses of sporadic CRC has shown that EOCRC is very unlikely to have microsatellite instability, BRAF mutations, or be a result of hypermethylation (CIMP+), all features of right-sided tumours,30,31 and characteristic of CMS1 subtype32.Therefore, changes in the tumour microbiome could contribute to the different patterns of clinical and molecular characteristics, including CMS subtypes, seen in EOCRC compared to LOCRC. Or alternatively the CMS status may favour microbial populations over another. While the EOCRC tumours in this study were predominantly CMS2/3, no associations were seen between these CRC-associated microbes and CMS type, with the numbers of patients in each CMS group being too small to integrate this data.
We found that EOCRC samples tended to have reduced microbial alpha diversity, with significantly lower species richness, when compared to tumours from the older patient group. This is consistent with research published by Xu et al, which also found decreased alpha diversity in the microbiome in early-onset tumour samples in their cohort.33 Reduced alpha diversity has been associated with several other disease states, including inflammatory bowel disease,34 irritable bowel syndrome,35 obesity,36 and even anxiety and stress.37 There are many factors that may contribute to the reduction of alpha diversity in the gut microbiome, including low-fibre diets, western or more industrialised lifestyles, caesarean section rates, and antibiotic usage.38,39,40 Many of these factors have increased in prevalence in recent decades, and therefore may be contributing to rising EOCRC incidence, however, whether a link between reduced alpha diversity in the gut corresponds to reduced alpha diversity in colorectal tumours remains to be investigated.
The trends of increasing incidence of EOCRC are occurring independently of CRC trends in older patients, and may be driven by changing environmental exposures that are disproportionately impacting younger patients. One possible candidate, sugary drink intake, has been increasing in younger people since the 1980’s,41 and has been shown to impact the microbiome,42 and has been shown to be an independent risk factor for EOCRC.43 This is also the case for obesity,44 antibiotic usage,45,46 Western-style diet,47 and even exposure to microplastics.48 The impact of environmental exposures on any individual’s microbiome will be different as people age. Microbial diversity is found to increase dramatically in early years, from infancy to adolescence, becoming more stable during adult life.49 Consequently, the impact of various environmental exposures on the gut microbiome is less pronounced, with older people displaying more established and resilient microbiological communities when compared with younger people.50 This may mean that changing environmental exposures disproportionately impact young people and provide a potential explanation for increasing EOCRC incidence, without corresponding increases in older adults.2
Our study also identified several bacterial phyla, genera and species that were more enriched in tumours from our younger cohort compared to the older patients, further supporting the theory that there are distinct tumour microbiomes in EOCRC when compared to LOCRC, but the exact mechanisms through which these differences may be driving carcinogenesis remain unclear. Interestingly, we did not find any significant differences in abundance of species already known to be associated with CRC development, such as, Bacteroides fragilis, Bacteroides thetaiotaomicron, Porphyromonas assacharolytica, Parvimonas micra and Prevotella intermedia, with the exception of Fusobacterium nucleatum.
By investigating the characteristics of those bacteria that were enriched, and linking them to molecular pathways that drive oncogenesis, there may be some clues as to how certain bacteria are driving EOCRC development. One genus found to be enriched in our EOCRC cohort was Anabaena. This is a genus of cyanobacteria, known for its capacity to produce toxins, namely anatoxin and microcystin.51 Microcystin is a hepatotoxic compound, classed as a carcinogen, and shown to induce expression of MYC,52 an oncogene found to be over-expressed in 70-80% of CRC.53 This may be particularly relevant to EOCRC given that MYC promotes tumour formation by inducing chromosomal instability, which is the most common oncogenic pathway in EOCRC.30,31 Microcystin has also been shown to activate the PI3K/Akt/Wnt/ß-catenin pathway, another chromosomal instability pathway commonly associated with EOCRC.54 Microcystin can be a contaminant in drinking water and studies have suggested an association between microcystin in drinking water and rates of CRC.55 Interestingly, microcystin has been shown to be the most abundant class of cyanotoxin found in Aotearoa New Zealand water bodies,56 and with blooms of cyanobacteria dependant on warmer temperatures57, their concentrations in drinking water may be increasing as a result of global warming, highlighting an important area for further research. Another genus enriched in our EOCRC tumour microbiomes was Schaalia, which was found by Xu et al33 to be a key microbe in their EOCRC tumour microbiome cohort. There is some evidence that changes in the enteric environment and immune activation caused by Schaalia metabolism may aggravate the injuries caused by inflammation,58 although further research into the exact mechanism of this association is required.
Another species of interest is Klebsiella variicola, found to be significantly enriched in our EOCRC cohort. Klebsiella variicola is found, through a combination of mechanisms, to disrupt the integrity of the intestinal mucus layer and lead to inflammation of the colon, and, therefore, potentially increasing CRC risk.59 Prevalence of this bacteria may be increasing as Klebsiella varciicola can colonise the gut via the ‘mouth-gut axis’ in patients with oral periodontitis, a condition with increasing prevalence, internationally, partly due to the increasing sugar content of food;60,61 as mentioned above, consumption of sugary drinks has been increasing in recent decades and has been linked to risk of EOCRC.41,43 While the microbial associations detailed above give indications of potential exposome-related impacts on colorectal carcinogenesis, this study was retrospective and included a relatively small number of early-onset cases. Future studies should include larger cohorts with robust control groups and mechanistic studies to determine whether these findings are causative.
Finally and importantly, there are compelling data that show the status of the microbiome impacts the effectiveness of immunotherapy62,63 and that certain bacteria can metabolise cytotoxic drugs64 such as those employed in the management and care of patients with CRC. In view of the differences in microbiome in tumours from patients with early and late onset CRC, evaluating patient outcomes in response to treatment modalities based upon each of these cohorts in respect to the tumour-resident microbiome is warranted and should be included in future larger studies.
There are several limitations of this study, the most notable of which is the small number of patients in our EOCRC cohort (n = 19). This limits statistical power and makes it difficult to draw strong conclusions about the role of different species in driving CRC formation. There are also limitations inherent to sampling the tumour microbiome. The composition of the microbiome in the tumour may be different from the bacteria present in normal mucosa, in the stool, or sampled via rectal swab.65 Likewise, we cannot establish whether these microbes are colonising due to the environment of the tumour, or if they have some causative role in tumour development. While the microbiome tends to remain stable throughout adulthood, it has been noted to change with advancing age. Future studies should include healthy age-matched controls to account for compositional changes due to aging.
There may also be temporal relationships between certain bacteria and different stages in colorectal cancer carcinogenesis, with the bacteria that drive early cancer development no longer being enriched in an established tumour. Examples of this include ‘passenger’ bacteria proposed in the ‘driver-passenger’ model. This is where carcinogenesis is triggered by colonisation of certain ‘driver’ bacteria that cause changes in the environment within the bowel allowing for colonisation by opportunistic ‘passenger’ bacteria, which can either promote or stall the development of CRC, and can potentially outcompete the original driver bacteria.66 Likewise, in the ‘keystone’ hypothesis, the presence of a specific bacteria, in low numbers may lead to certain key changes in the bowel that drive microbial dysbiosis, inflammation and subsequent carcinogenesis.67 Finally, the ‘hit and run’ hypothesis describes a situation where temporary colonisation by certain bacteria may be sufficient to trigger a carcinogenic chain of events, due to the action of specific toxins (e.g. CagA from Helicobacter pylori initiating the development of gastric adenocarcinoma), implying that the presence of these bacteria may initiate carcinogenesis but not be required for the subsequent tumour development, and may be absent or sparse in the cancer itself.68
This study adds to the growing body of evidence that EOCRC is a distinct disease from LOCRC. Changing environmental exposures and the impact that these have on patients’ microbiomes may be involved in the increased incidence of EOCRC.
Author contribution: OW carried out laboratory analysis and wrote the initial draft of the manuscript; AS carried out bioinformatics analysis and contributed towards writing of the manuscript; KW, RR and AH were responsible for sample collection, contributed to study design and interpretation of the data; RP and FF designed the study, contributed to manuscript revision and interpretation of data. All authors have read and approved the final manuscript draft.
Funding: RP was funded by the Health Research Council of New Zealand and Bowel Cancer Research Aotearoa.
Competing interests statement: The authors declare no competing interests.
Ethics, Consent to Participate: The study was approved by the University of Otago Human Ethics Committee (approval number: H16/037), the Health and Disability Ethics Committee, New Zealand (approval number: 18/STH/40/AM01), and the Human Research Ethics Committees of Australia (approval number: HREC 14/85). All participants in this cohort provided written, and informed consent.
Data availability: Sequencing data can be found under Bioproject ID PRJNA815861 and Bioproject ID PRJNA788974 in the NCBI SRA database.
References
-
Siegel RL, Soerjomataram I, Hayes RB, Bray F, Weber TK, Jemal A. Global patterns and trends in colorectal cancer incidence in young adults. Gut. 2019;68:2179-2185.
-
Waddell O, Pearson J, McCombie A, et al. The incidence of early onset colorectal cancer in Aotearoa New Zealand: 2000-2020. BMC Cancer. 2024;24(1):456.
-
Gandhi J, Davidson C, Hall C, et al. Population-based study demonstrating an increase in colorectal cancer in young patients. British Journal of Surgery. 2017;104(8):1063-1068.
-
Bailey CE, Hu CY, You YN, et al. Increasing disparities in age-related incidence of colon and rectal cancer in the United States, 1975-2010. JAMA Surg. 2015;150(1)
-
Abdelsattar ZM, Wong SL, Regenbogen SE, Jomaa DM, Hardiman KM, Hendren S. Colorectal cancer outcomes and treatment patterns in patients too young for average-risk screening. Cancer. 2016;122(6):929-934.
-
O’connell JB, Maggard MA, Liu JH, Etzioni DA, Livingston EH, Ko CY. Do Young Colon Cancer Patients Have Worse Outcomes? 2004.
-
Kolarich A, Thomas ;, George J, et al. Rectal Cancer Patients Younger Than 50 Years Lack a Survival Benefit From NCCN Guideline-Directed Treatment for Stage II and III Disease. Cancer. 2018;124:3510-3519.
-
Chittleborough TJ, Gutlic I, Pearson JF, et al. Increasing incidence of young-onset colorectal carcinoma A 3-country population analysis. Dis Colon Rectum. 2020;63(7):903-910.
-
Wild CP. Complementing the genome with an “exposome”: the outstanding challenge of environmental exposure measurement in molecular epidemiology. Cancer Epidemiol Biomarkers Prev. 2005;14(8):1847-1850.
-
Hofseth LJ, Hebert JR, Chanda A, et al. Early-onset colorectal cancer: initial clues and current views. Nat Rev Gastroenterol Hepatol. 2020;17(6):352-364.
-
Moon JY, Kye BH, Ko SH, Yoo RN. Sulfur Metabolism of the Gut Microbiome and Colorectal Cancer: The Threat to the Younger Generation. Nutrients 2023, Vol 15, Page 1966. 2023;15(8):1966.
-
Keenan JI, Frizelle FA. Toxigenic gut bacteria, diet and colon carcinogenesis. J R Soc N Z. 2020;50(3):418-433.
-
Louis P, Hold GL, Flint HJ. The gut microbiota, bacterial metabolites and colorectal cancer. Nat Rev Microbiol. 2014;12(10):661-672.
-
Purcell R V, Pearson J, Aitchison A, Dixon L, Frizelle FA, Keenan JI. Colonization with enterotoxigenic Bacteroides fragilis is associated with early-stage colorectal neoplasia. PLoS One. 2017;12(2).
-
Zhong L, Zhang X, Covasa M. Emerging roles of lactic acid bacteria in protection against colorectal cancer. World Journal of Gastroenterology : WJG. 2014;20(24):7878.
-
Faghfoori Z, Faghfoori MH, Saber A, Izadi A, Yari Khosroushahi A. Anticancer effects of bifidobacteria on colon cancer cell lines. Cancer Cell Int. 2021;21(1):1-12.
-
Kong C, Liang L, Liu G, et al. Integrated metagenomic and metabolomic analysis reveals distinct gut-microbiome-derived phenotypes in early-onset colorectal cancer. Gut. 2023;72(6):1129-1142.
-
Barot S V., Sangwan N, Nair KG, et al. Distinct intratumoral microbiome of young-onset and average-onset colorectal cancer. EBioMedicine. 2024;100.
-
Purcell R V., Schmeier S, Lau YC, Pearson JF, Frizelle FA. Molecular subtyping improves prognostication of Stage 2 colorectal cancer. BMC Cancer. 2019;19(1):1-9.
-
Sulit AK, Wilson K, Pearson J, et al. Human gene and microbial analyses in rectal cancer complete responses to radiotherapy. BJS Open. 2023;7(3).
-
Purcell R V., Visnovska M, Biggs PJ, Schmeier S, Frizelle FA. Distinct gut microbiome patterns associate with consensus molecular subtypes of colorectal cancer. Scientific Reports 2017 7:1. 2017;7(1):1-12.
-
Sulit AK, Daigneault M, Allen-Vercoe E, et al. Bacterial lipopolysaccharide modulates immune response in the colorectal tumor microenvironment. npj Biofilms and Microbiomes 2023 9:1. 2023;9(1):1-8.
-
Sulit AK, Wilson K, Pearson J, et al. Human gene and microbial analyses in rectal cancer complete responses to radiotherapy. BJS Open. 2023;7(3).
-
Sulit AK, Kolisnik T, Frizelle FA, Purcell R, Schmeier S. MetaFunc: taxonomic and functional analyses of high throughput sequencing for microbiomes. Gut Microbiome. 2023;4:e4.
-
Menzel P, Ng KL, Krogh A. Fast and sensitive taxonomic classification for metagenomics with Kaiju. Nat Commun. 2016;7.
-
McMurdie PJ, Holmes S. phyloseq: An R Package for Reproducible Interactive Analysis and Graphics of Microbiome Census Data. PLoS One. 2013;8(4):e61217.
-
Love, M.I., W. Huber, and S. Anders, Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol, 2014. 15(12): p. 550.
-
Risso, D., et al., A general and flexible method for signal extraction from single-cell RNA-seq data. Nat Commun, 2018. 9(1): p. 284.
-
Flynn KJ, Ruffin MT, Kim Turgeon D, Schloss PD. Spatial variation of the native colon microbiota in healthy adults. Cancer Prevention Research. 2018;11(7):393-401.
-
Kirzin S, Marisa L, Guimbaud R, et al. Sporadic Early-Onset Colorectal Cancer Is a Specific Sub-Type of Cancer: A Morphological, Molecular and Genetics Study. PLoS One. 2014;9(8):e103159.
-
Chouhan H, Ferrandon S, Devecchio J, Kalady MF, Church JM. A Changing Spectrum of Colorectal Cancer Biology with Age: Implications for the Young Patient. Dis Colon Rectum. 2019;62(1):21-26.
-
Marx O, Mankarious M, Yochum G. Molecular genetics of early-onset colorectal cancer. World J Biol Chem. 2023;14(2):13.
-
Xu Z, Lv Z, Chen F, et al. Dysbiosis of human tumor microbiome and aberrant residence of Actinomyces in tumor-associated fibroblasts in young-onset colorectal cancer. Front Immunol. 2022;13:1008975.
-
Alam MT, Amos GCA, Murphy ARJ, Murch S, Wellington EMH, Arasaradnam RP. Microbial imbalance in inflammatory bowel disease patients at different taxonomic levels. Gut Pathog. 2020 Jan 4;12:1.
-
Carroll IM, Ringel-Kulka T, Siddle JP, Ringel Y. Alterations in composition and diversity of the intestinal microbiota in patients with diarrhea-predominant irritable bowel syndrome. Neurogastroenterology & Motility. 2012;24(6):521-e248.
-
Pinart M, Dötsch A, Schlicht K, et al. Gut microbiome composition in obese and non-obese persons: A systematic review and meta-analysis. Nutrients. 2022;14(1).
-
Johnson KVA. Gut microbiome composition and diversity are related to human personality traits. Hum Microb J. 2020;15:100069.
-
Jakobsson HE, Abrahamsson TR, Jenmalm MC, et al. Decreased gut microbiota diversity, delayed Bacteroidetes colonisation and reduced Th1 responses in infants delivered by Caesarean section. Gut. 2014;63(4):559-566.
-
Mosca A, Leclerc M, Hugot JP. Gut microbiota diversity and human diseases: Should we reintroduce key predators in our ecosystem? Front Microbiol. 2016;7(MAR):178205.
-
Yatsunenko T, Rey FE, Manary MJ, et al. Human gut microbiome viewed across age and geography. Nature. 2012;486(7402):222.
-
Wang YC, Bleich SN, Gortmaker SL. Increasing caloric contribution from sugar-sweetened beverages and 100% fruit juices among US children and adolescents, 1988-2004. Pediatrics. 2008;121(6).
-
Satokari R. High Intake of Sugar and the Balance between Pro- and Anti-Inflammatory Gut Bacteria. Nutrients 2020, Vol 12, Page 1348. 2020;12(5):1348.
-
Hur J, Otegbeye E, Joh HK, et al. Sugar-sweetened beverage intake in adulthood and adolescence and risk of early-onset colorectal cancer among women. Gut. 2021;70(12):2330-2336.
-
Sanford NN, Giovannucci EL, Ahn C, Dee EC, Mahal BA. Obesity and younger versus older onset colorectal cancer in the United States, 1998–2017. J Gastrointest Oncol. 2020;11(1):121.
-
Dik VK, van Oijen MGH, Smeets HM, Siersema PD. Frequent Use of Antibiotics Is Associated with Colorectal Cancer Risk: Results of a Nested Case–Control Study. Dig Dis Sci. 2016;61(1):255-264.
-
Jiang F, Boakye D, Sun J, et al. Association between antibiotic use during early life and early-onset colorectal cancer risk overall and according to polygenic risk and FUT2 genotypes. Int J Cancer. Published online 2023.
-
Moon JY, Kye BH, Ko SH, Yoo RN. Sulfur Metabolism of the Gut Microbiome and Colorectal Cancer: The Threat to the Younger Generation. Nutrients 2023, Vol 15, Page 1966. 2023;15(8):1966.
-
Li S, Keenan JI, Shaw IC, Frizelle FA. Could Microplastics Be a Driver for Early Onset Colorectal Cancer? Cancers 2023, Vol 15, Page 3323. 2023;15(13):3323.
-
Odamaki T, Kato K, Sugahara H, et al. Age-related changes in gut microbiota composition from newborn to centenarian: A cross-sectional study. BMC Microbiol. 2016;16(1):1-12.
-
Hildebrand F, Gossmann TI, Frioux C, et al. Dispersal strategies shape persistence and evolution of human gut bacteria. Cell Host Microbe. 2021;29(7):1167-1176.e9.
-
Carmichael WW. Cyanobacteria secondary metabolites—the cyanotoxins. Journal of Applied Bacteriology. 1992;72(6):445-459.
-
Fan H, Cai Y, Xie P, et al. Microcystin-LR stabilizes c-myc protein by inhibiting protein phosphatase 2A in HEK293 cells. Toxicology. 2014;319(1):69-74.
-
Erisman MD, Rothberg PG, Diehl RE, Morse CC, Spandorfer JM, Astrin SM. Deregulation of c-myc Gene Expression in Human Colon Carcinoma Is Not Accompanied by Amplification or Rearrangement of the Gene. Mol Cell Biol. 1985;5(8):1969-1976.
-
Tang Y, Yi X, Zhang X, et al. Microcystin-leucine arginine promotes colorectal cancer cell proliferation by activating the PI3K/Akt/Wnt/β-catenin pathway. Oncol Rep. 2023;49(1).
-
Zhu L, Hai Y, Kun C. Relationship between microcystin in drinking water and colorectal cancer. Biomed Environ Sci. 2002;15(2).
-
Wood SA, Holland PT, Stirling DJ, et al. Survey of cyanotoxins in New Zealand water bodies between 2001 and 2004. N Z J Mar Freshwater Res. 2006;40(4):585-597.
-
Paerl HW, Huisman J. Climate: Blooms like it hot. Science (1979). 2008;320(5872):57-58.
-
Li J, Li Y, Zhou Y, Wang C, Wu B, Wan J. Actinomyces and Alimentary Tract Diseases: A Review of Its Biological Functions and Pathology. Biomed Res Int. 2018;2018.
-
Gong J, Yu J, Yin S, et al. Mesenteric Adipose Tissue-Derived Klebsiella variicola Disrupts Intestinal Barrier and Promotes Colitis by Type VI Secretion System. Advanced Science. 2023;10(12):2205272.
-
World Health Organisation (WHO). Oral health. 2023.
-
Kitamoto S, Nagao-Kitamoto H, Jiao Y, et al. The Intermucosal Connection between the Mouth and Gut in Commensal Pathobiont-Driven Colitis. Cell. 2020;182(2):447-462.e14.
-
Li X, Zhang S, Guo G, Han J, Yu J. Gut microbiome in modulating immune checkpoint inhibitors. EBioMedicine. 2022;82.
-
Lu Y, Yuan X, Wang M, et al. Gut microbiota influence immunotherapy responses: mechanisms and therapeutic strategies. Journal of Hematology & Oncology 2022 15:1. 2022;15(1):1-20.
-
Huang J, Liu W, Kang W, et al. Effects of microbiota on anticancer drugs: Current knowledge and potential applications. EBioMedicine. 2022;83.
-
Sun S, Zhu X, Huang X, et al. On the robustness of inference of association with the gut microbiota in stool, rectal swab and mucosal tissue samples. Scientific Reports |. 123AD;11:14828.
-
Tjalsma H, Boleij A, Marchesi JR, Dutilh BE. A bacterial driver-passenger model for colorectal cancer: beyond the usual suspects. Nat Rev Microbiol. 2012;10(8):575-582.
-
Hajishengallis G, Darveau RP, Curtis MA. The keystone-pathogen hypothesis. Nat Rev Microbiol. 2012;10(10):717-725.
-
Hatakeyama M. Helicobacter pylori CagA and gastric cancer: a paradigm for hit-and-run carcinogenesis. Cell Host Microbe. 2014;15(3):306-316.





